What if you could cultivate your own gut microbiome — one fibre, one probiotic, one antibiotic at a time? GutQuest is a free, open-source serious game that turns the living ecosystem of your intestinal microbiota into an interactive 2D simulation. No arcade reflexes required — just curiosity, ecological thinking, and a willingness to experiment with the 38 trillion microbial citizens inside you.
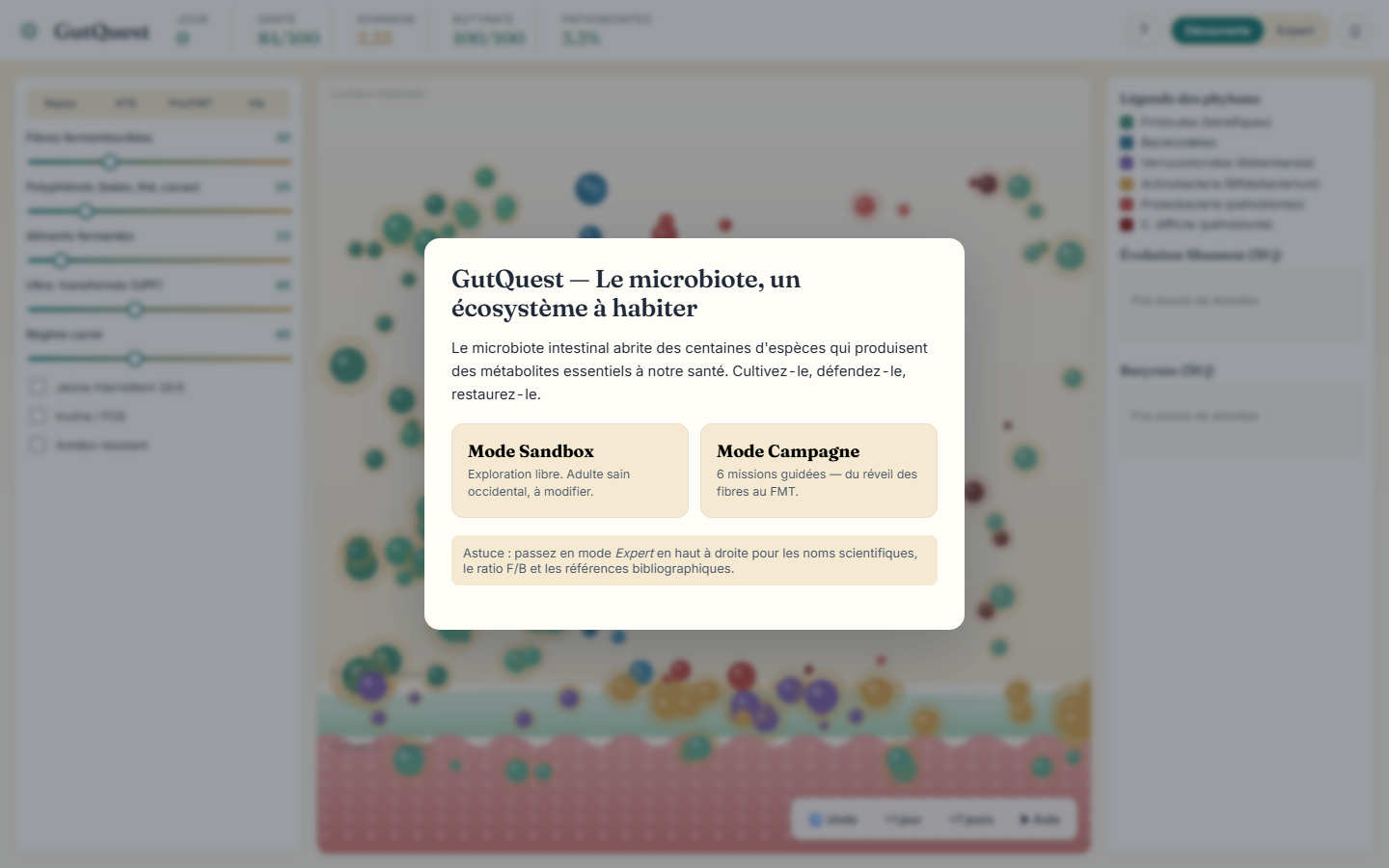
The pitch: an ecological simulation, not an arcade game
GutQuest models the human gut as what it genuinely is: a competitive, cooperative, dynamic ecosystem shaped by diet, drugs, lifestyle, and time. The simulation tracks twelve microbial species and genera, each with its own substrate preferences, metabolic outputs, preferred anatomical niche, and ecological role. This is not a simplified bar chart — it is a population-dynamics engine inspired by ecological modelling, rendered as an animated 2D cross-section of the intestinal wall.
The twelve microbial players are distributed across three anatomical niches:
| Niche | Key species | Ecological role |
|---|---|---|
| Intestinal lumen | Faecalibacterium prausnitzii, Roseburia intestinalis, Eubacterium rectale, Bacteroides, Prevotella, commensal E. coli, Enterococcus, C. difficile, Klebsiella | Fermentation, SCFA production, pathobiont colonisation |
| Mucus layer | Akkermansia muciniphila, Bifidobacterium | Barrier maintenance, mucoprotection |
| Crypts / epithelium | Lactobacillus | Immune priming, colonisation resistance |
Each species is assigned a phylum (Firmicutes, Bacteroidetes, Verrucomicrobia, Actinobacteria, Proteobacteria), a role (beneficial keystone, commensal, or pathobiont), and a set of substrates it metabolises — fermentable fibres, resistant starch, mucins, simple sugars — alongside the metabolites it produces: butyrate, propionate, acetate, lactate, or bacterial toxins.
Population dynamics run every simulated day using a logistic growth equation modulated by substrate availability, niche fit, inter-species competition, intervention multipliers, and a stress coefficient. Total carrying capacity is conserved: when beneficials collapse, pathobionts fill the vacated niches — exactly as documented in the antibiotic-disruption literature (Dethlefsen & Relman, 2011).
How it works: the intervention toolkit
The left-hand panel gives you four categories of intervention. Each one translates directly into ecological consequences visible in the animated canvas.
Diet
Dietary toggles and sliders modulate the substrate pool available in the lumen. Adding fermentable fibres (legumes, oats, vegetables) triggers a bloom of F. prausnitzii, Roseburia, and Eubacterium rectale — the main butyrate producers. Polyphenols (berries, green tea, extra-virgin olive oil) selectively boost Akkermansia and Bifidobacterium, thickening the mucus layer on screen. Fermented foods (yoghurt, kefir, sauerkraut, kimchi) provide a modest colonisation boost to Lactobacillus.
On the other side of the equation: ultra-processed foods (UPF) crash fibre availability, spike simple sugar levels, and trigger a visible expansion of Enterobacteriaceae while the mucus layer thins. Intermittent fasting (16:8) creates lumenal substrate-free windows during which Akkermansia grazes on mucins — a counterintuitive benefit modelled from experimental data. Prebiotics — inulin, FOS, GOS, resistant starch — act as precision fertilisers for specific target populations.
Antibiotics
The antibiotic module lets you choose spectrum (narrow vs. broad), duration (3, 7, or 14 days), and molecule: amoxicillin, amoxicillin-clavulanate, ciprofloxacin, clindamycin, oral vancomycin, or metronidazole. The consequences are immediate and brutal. Broad-spectrum courses collapse diversity by 50–95 %, opening a classic colonisation window for C. difficile and Klebsiella. Clindamycin and amoxicillin-clavulanate carry the highest C. difficile risk — exactly as in clinical practice. Post-antibiotic resilience is partial: diversity recovers over 1–3 months with a fibre-rich diet, but never fully reconstitutes without active intervention (Dethlefsen & Relman, 2011).
Probiotics, prebiotics & FMT
Lactobacillus/Bifidobacterium probiotics provide a transient boost (days 0–14) with limited long-term re-implantation — consistent with clinical trial data. Saccharomyces boulardii offers targeted protection during antibiotic courses. Exogenous butyrate (postbiotic) rapidly restores barrier integrity in the short term. Faecal Microbiota Transplantation (FMT) is modelled as a hard reset — it dramatically restores diversity and is the only intervention that reliably clears recurrent C. difficile (Cammarota et al., 2017 European consensus). On screen, FMT triggers the most visually dramatic shift: the colony landscape rebuilds itself within a few simulated days.
Lifestyle
Chronic stress (elevated cortisol) suppresses Lactobacillus and increases gut permeability. Sleep restriction below 6 hours disrupts circadian entrainment of the microbiota, reducing diversity. Regular moderate exercise independently increases Akkermansia and butyrate production. A tropical travel scenario exposes the ecosystem to ETEC E. coli and antibiotic-resistant organism colonisation — testing the player’s prophylaxis and recovery strategy.
The health dashboard: five indicators + one global score
Every simulated day, the HUD updates five composite indicators:
- Shannon diversity (α-diversity H′) — displayed 0–4; the ecological gold standard for microbial richness and evenness
- Butyrate production — weighted sum of outputs from the three main butyrate producers; drives barrier health
- Barrier integrity — a function of Akkermansia, Bifidobacterium, and butyrate levels; visible as mucus thickness on screen
- Pathobiont load — percentage of total bacterial population occupied by C. difficile, Klebsiella, and dysbiotic Enterococcus
- Global Microbiome Health Score (0–100) — a composite: 25 % diversity + 20 % butyrate + 20 % barrier + 15 % (1 − pathobionts) + 10 % richness + 10 % Firmicutes/Bacteroidetes balance
Two reading levels: one game for everyone
A toggle in the top-right corner switches between Discovery mode and Expert mode — persistent throughout the session.
Discovery mode
Species are labelled with plain-language roles (« good bacteria », « protective bacteria », « opportunistic bacteria »). Tooltip bubbles explain each intervention in everyday terms. A built-in glossary defines unfamiliar words. The Health Score is front and centre. Designed for curious patients, students, and informed members of the public — no prior microbiology required.
Expert mode
Full scientific nomenclature throughout. The HUD displays the raw Shannon H′ index and the Firmicutes/Bacteroidetes ratio. Tooltips cite primary literature: Sokol et al., 2008 on F. prausnitzii as an anti-inflammatory keystone in Crohn’s disease; Cammarota et al., 2017 on the European FMT consensus; Wu et al., 2011 on Bacteroides vs. Prevotella enterotypes; David et al., 2014 on diet changing the microbiome within 24 hours. Designed for GPs, gastroenterologists, dietitians, pharmacists, and undergraduate / CME-level educators.
Six clinical missions
Beyond the sandbox, GutQuest offers a six-mission campaign, each grounded in a realistic clinical or lifestyle scenario:
- The Fibre Awakening (Discovery) — Starting from a fibre-depleted, UPF-dominated microbiome, the player must push butyrate above 60/100 within 30 simulated days. Core lesson: fermentable fibres → short-chain fatty acids → barrier integrity.
- Antibiotics Under Control — A patient requires antibiotic treatment for pneumonia. The player chooses the molecule, spectrum, and duration, then selects adjuvant strategies. Objective: cure the infection and keep Shannon diversity above 2.5. Core lesson: rational antibiotic use, Saccharomyces boulardii protection.
- Saving Mr D. from C. difficile (Expert) — After three antibiotic courses, the patient has recurrent Clostridioides difficile colitis. Options: extended oral vancomycin, fidaxomicin, or FMT. Objective: eradicate C. difficile and restore Shannon above 3. Core lesson: FMT indications, niche ecology, colonisation resistance.
- The Marathon Runner — Optimising the microbiome of an endurance athlete preparing a marathon. Focus on Akkermansia, Veillonella, and butyrate for performance and recovery. Core lesson: the microbiome–exercise axis.
- Tropical Travel — Managing traveller’s diarrhoea risk, ETEC colonisation, and antibiotic-resistant organism exposure. Core lesson: travel medicine, decolonisation strategies, microbiome resilience.
- Reset After Burnout (Expert) — Two years of chronic stress, degraded diet, and less than five hours of sleep per night. A 90-day multimodal recovery plan targeting the gut–brain axis. Core lesson: the multifactorial nature of dysbiosis, the microbiota–gut–brain dialogue.
For healthcare professionals
GutQuest is designed as an educational companion tool for clinical settings. Use it in consultation to visualise why antibiotic choice and duration matter, why fibre diversity outperforms single-strain probiotics, or why FMT produces results that no other intervention can replicate. The dual reading level makes it equally suitable for patient discussions and for undergraduate teaching or CME modules in gastroenterology, infectious disease, nutrition, and pharmacy.
Important: GutQuest is a didactic simulation tool. It is not a medical device, does not constitute medical advice, and should not be used to guide individual clinical decisions. Parameters are calibrated from published literature but are deliberately simplified for educational clarity.
Key takeaways
- Simplified but scientifically coherent — every mechanism is grounded in peer-reviewed ecology; parameters are derived from published studies, then scaled for interactivity.
- 100 % open-source — the full codebase is published under an open licence on GitHub; contributions, forks, and bug reports are welcome.
- Hosted on GitHub Pages — no installation, no account, no data collection. Accessible from any modern browser.
- Desktop and mobile compatible — the responsive layout adapts to tablets and smartphones; opens in a new tab for an immersive fullscreen experience.
- Completely free — no paywall, no premium tier, no advertisement. Open science, open education.
Mini French → English glossary for in-game terms
The game interface is in French. Here is a quick reference for every term you will encounter on screen:
| French (in-game) | English equivalent | Context |
|---|---|---|
| Lumière | Lumen | Central cavity of the intestine; richest niche for fermentation |
| Mucus | Mucus layer | Protective gel secreted by goblet cells; thickness = barrier health |
| Cryptes | Crypts | Epithelial invaginations near the intestinal wall; microaerobic niche |
| Fibres | Fibres / Dietary fibre | Fermentable substrates for beneficial bacteria |
| Antibiotique / ATB | Antibiotic | Bacterial population disruptor; spectrum and duration are key |
| Probiotique | Probiotic | Live microbial supplement; transient benefit modelled J0–J14 |
| Prébiotique | Prebiotic | Selectively fermented substrate (inulin, FOS, GOS) |
| Greffe fécale / FMT | Faecal Microbiota Transplantation (FMT) | Ecosystem reset; first-line for recurrent C. difficile |
| Diversité | Diversity | Shannon H′ index: ecological richness + evenness (0–4) |
| Butyrate | Butyrate | Key short-chain fatty acid; fuels colonocytes, seals the barrier |
| Barrière | Barrier integrity | Composite score of mucus thickness and tight junction health |
| Pathobionte | Pathobiont | Normally commensal species that becomes harmful under dysbiosis |
| Santé | Health Score | Global composite score 0–100 |
| Jour | Day | Simulation time unit; advance 1 day, 7 days, or auto-play |
Limitations and scientific honesty
GutQuest is a didactic model, not a digital twin of your gut. Several deliberate simplifications are worth naming explicitly:
- Twelve species cannot capture 500. The human gut hosts hundreds of species and thousands of strains. GutQuest selects twelve representatives chosen for their educational relevance, metabolic diversity, and clinical significance — not for exhaustive coverage.
- Parameters are calibrated from the literature, but compressed. Growth rates, substrate preferences, and intervention multipliers are derived from published data (see references cited in Expert mode) but are scaled and simplified to produce instructive, readable dynamics within a game session.
- Individual variability is not modelled. The real microbiome is shaped by genetics, age, birth mode, early colonisation history, and geographic origin — none of which appear in GutQuest. The simulation represents a stylised « average Western adult » baseline.
- This is not a microbiome prescription. No simulation output should be interpreted as a personalised dietary or therapeutic recommendation. Decisions about antibiotics, probiotics, or FMT require clinical assessment.
- No data is stored. The game runs entirely in-browser with no persistent storage, no account, and no tracking.
Open source — contribute and build on it
The entire GutQuest codebase — population model, rendering engine, mission scripts, and UI — is open at github.com/JETPOD/gutquest. It is built with HTML5 Canvas 2D and vanilla JavaScript (ES modules, no framework dependency), making it straightforward to fork, adapt, or extend. Google Fonts (Fraunces + Inter) are the only external dependency. Pull requests, issue reports, translations, and educational forks are all welcome.
If you are a researcher, educator, or developer interested in extending the ecological model, adding new species, building a localised version, or integrating GutQuest into a learning management system, the repository is your starting point.
🇫🇷 Lire cet article en français : GutQuest : explorez votre microbiote intestinal comme un écosystème vivant
NutriCellScience, Mark DOWN — EN edition
Laisser un commentaire